|
See also: Topics and links to materials on the topics to be discussed at the Davis Genealogy Club panel discussion on April 16, 2019.
This was written in 2007 when the most common DNA tests were with Mitochondrial DNA for women and y-DNA for men. Where does your DNA come from For Genetic Genealogy, which is the application of DNA testing to genealogy research, three types of DNA can provide information useful in conjunction with genealogy research. These types are the Y chromosome (Y-DNA), Mitochondrial (mtDNA) and autosomal (auDNA). y-DNA - The Y-chromosome only exists in males, so is used for tracing male lines. Mitochondrial DNA (mtDNA), is passed only in the egg outside the cell nucleus, so comes from your Mother and is useful for tracing female lines. Autosomes are all chromosomes other than the X and Y. They can be used to link you to cousins on both parents side. Within the non-coding (or Junk) DNA regions (regions which are not part of genes. 95% or more of the DNA) are certain known locations (loci) where a short segment of DNA (2-5 base-pairs) will stutter, repeating itself a number of times. This is known as a Short Tandem Repeat (STR). There are over 100 of these STRs called markers. They are given DNA Y-chromosome Segment (DYS) numbers to identify them. The number of repeats (Allele value) changes (mutates) over time. Testing involves counting the number of repeats for a group of these Testing labs typically use 12, 25, 37 or 67 marker tests to do for genealogy. The 25 marker test it the most common. These STRs have a couple of properties that make them attractive for genealogy.
mtDNA testing looks for single nucleotide (base-pair) polymorphisms (SNPs) in certain regions.
Testing:
DNA Overview: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
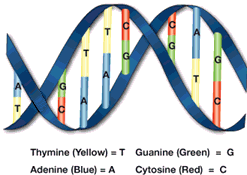 |
The rungs of the double helix DNA molecule are what vary from one person to another. They are made up of pairs of four bases, Guanine (green) always pairs with cytosine (red) and thymine (yellow) always pairs with adenine (blue).
The order of these bases is called the DNA sequence. You can write the sequence by listing the bases on either side. A STR has pattern of 2-5 bases that repeats. DYS # 393 has the following repeat: (GATA)(GATA)(GATA)(GATA)(GATA)(GATA)(GATA) ... 70% of the population has 13 repeats, 12% have 12, 12% have 14, the rest having 10, 11, 15 or 16 repeats. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
A sample Haplotype from a 25 marker test:
Anthropologists break down the Y-chromosome into branches called Haplogroups or clades, which distinguish major branches of Homo Sapiens. Haplogroup R1b1 is the most common haplogroup in European populations. It is believed to have expanded throughout Europe as humans re-colonized after the last glacial maximum 10-12 thousand years ago.
Migration charts show how verious groups migrated out of Africa.
See: Y-Chromosome Nomenclature System at U. Arizona
for a report by the Y-Chromosome Consortium (YCC) where they say:
An article "DNA testing for Genealogy" at GenUKi.org.uk says: Companies:
Terms: Allele - The number of repeats of a sequence in a marker Distance - A relative indication of how closely you are related to someone. DYS - DNA Y-chromosome Segment - See marker haplotype - A string of allelle values for different markers Haplogroups - Identifies the person's major population group and provides information about the ancient origin. R1b1 is the most common haplogroup in European populations. Locus - Pysical locaation of a marker marker - Repeated base-pair sequences at specific locations along the Y chromosome. They are given DYS numbers to distinguish them. non-coding DNA - DNA (sometimes called junk or filler DNA) which does not contain instructions for making proteins (or other cell products such as noncoding RNAs) This accounts for 95-98% of the DNA SNP - single nucleotide polymorphism - A change in your Y-DNA sequence at a location other than those in the 12, 25, 37, and 67 marker tests. SNPS are unique to specific haplogroups so SNP tests such as the Backbone and Deep SNP tests are used to identify haplogroups and their subclades respectively. STR - Short Tandem Repeat - Short markers with 8-36 repeats of 2-5 base-pair sequences (DNA codes). These sequences are sometimes referred to as genes but they are not the same as the genes we think, which are much longer and used in coding for creating proteins determining amongst other things physical characteristics. TMRCA - Time to the Most Recent Common Ancestor. Y-STR - STR's on the Y-chromosome Average of 1.3% nonpaternity events per generation Rate of mutation = .002-.003 per marker. In other words, we could expect any marker to mutate once in 300-500 generations.
See: Books: "Trace Your Roots with DNA: Use Your DNA to Complete Your Family Tree:", Megan Smolenyak, Ann Turner Adam's Curse, Bryan Sykes Book List at FamilyTreeDNA
Links:
Return to biology
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||